|
PinkMonkey Online Study Guide-Biology
vi. Complementary base pairing : In each pair,
the two bases of the opposite strands are joined by hydrogen bonds. A
and T are joined by two hydrogen bonds, while G and C are joined by three
hydrogen bonds. This is called complementary base pairing. The two strands
are thus held together all along their lengths by these hydrogen bonds.
v. Purine : Pyrimidine ratio : Because of the fixed or complementary base pairing in the DNA molecule, the total number of A is equal to the total number of T and the total number of G is equal to the total number of C. In other words, (A+G)= (T+C). Hence, purines: pyrimidines ratio is 1:1.
vi. C-3 and C-5 ends of the strand : In each strand
one end of the strand has one free phosphate group on carbon-5 of the
sugar molecule. This is the end of the strand is called C-5 (or 5') end.
The other end of the strand has a free -OH on carbon-3 of the sugar molecule.
This is called C-3 (or 3') end of the strand (Figure 8.4c).
vii. Antiparallel nature of strands: The two strands
are oppositely oriented and hence are called antiparallel. This means,
the 3' end of one strand is adjacent to the 5' end of the other strand
(Fig. 8.4c) . This is because, the phosphate-sugar linkages run in opposite
directions in the two strands.
viii. Dimensions: The diameter of the DNA double
helix is 20 Ao. The length of each
complete spiral (turn or pitch) of the molecule measures 34 Ao.
10 pairs of nucleotides are present in each complete spiral. Therefore,
each nucleotide in the strand occupies a distance of 3.4A0.
(c) Circular DNA molecules : Chromosomes of most
prokaryotes (e.g. bacteria, cyanobacteria,) are circular molecules of
DNA.
In bacteria, there is no organized nucleus. The bacterial
nucleoid consists of a single circular DNA molecule (bacterial
chromosome). The molecule has two complementary strands forming a covalently
closed circle. Generally, the circular molecule is present in a highly
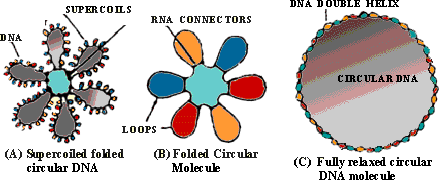
folded and suspercoiled state (Figure 8.5A). This is expected because
the diameter of a bacterial cell (e.g. Escherichia coli) is about
1-2 microns while the total length of the circular DNA is about 1100 microns.
The circular molecule has 40-50 folds or looped domains. These
folds are held in position by RNA molecules (RNA connectors) and some
non-histone proteins associated with the bacterial chromosome. (Histones
are absent in bacteria). The DNA segment in each loop is supercoiled independently.
Because of this characteristic formation of loops as well as the supercoils
within the loops, the large circular DNA molecule can be packed into a
small bacterial cell. Otherwise, the relaxed and fully expanded
circular molecule (Figure 8.5) would be too large (about 350 microns
diameter) to be contained in the bacterial cell. The coils of the supercoiled
circle can be relaxed by treatment with enzymes such as RNAse or DNAse.
The uncoiling occurs due to a break (nick) in one or both the strands
of DNA.
In some viruses, e.g. certain bacteriophages, the circular
DNA is single stranded. It becomes double stranded only during replication
(replicatable form).
 Click here to enlarge
Click here to enlarge
Figure 8.5
|